plot07.m - Overlaying a sine function on noisy data
period = 10; % define the period of the signal t = linspace(0,period,200); % 200 data points evenly spaced over 1 period % create a sine wave and add Gaussian noise to it % plot as blue data points y2 = sin(2*pi/period*t) + 0.5*randn(1,length(t)); plot(t,y2,'bo', 'MarkerSize',2, 'MarkerFaceColor', 'b'); % prevent future plots from overwriting current plot hold all % create a sine wave with no noise and plot as red curve y = sin(2*pi/period*t); plot(t,y,'r-') % red line % label axes xlabel('t') ylabel('y(t)') axis([0 period -4 4]); legend('sine + noise','sine');
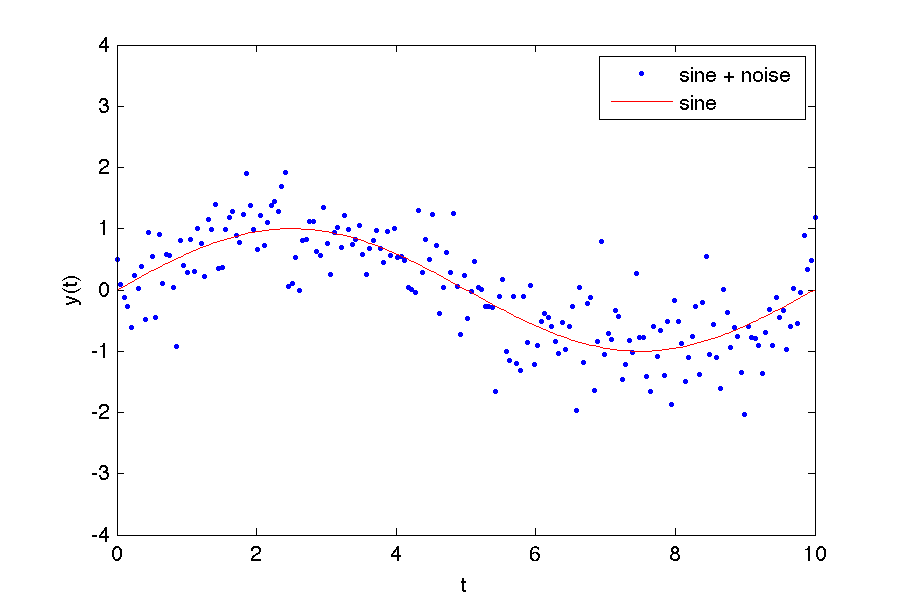
This code is an example of a common problem in experimental physics. You want to overlay a theoretical model curve on some noisy data to see if your model matches the data. In this case, we 'faked' a data set by just added Gaussian noise to a sine wave.
The noise is created with the randn() command. It generates an array of random values drawn from a Gaussian distribution with mean = 0 and variance = 1.
We use the hold all command to prevent the second plot from overwriting the first.
